Epithelial Cell Migration and Transformation
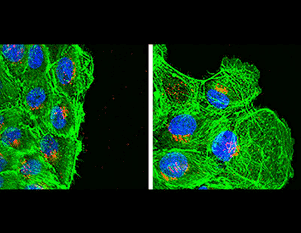
Focal adhesion dynamics in migrating MCF10A cells in which CRL5 activity has been inhibited. MCF10A cells expressing EYFP-vinculin were treated with siRNA against Cul5. Cells were removed from the upper part of the slide and the remaining cells are migrating into the wound. Cells were imaged every 2 min on a 100x TIRF objective, starting approximately 6 hr after wounding. Length of movie 120 min. DOI: 10.7554/eLife.17440.007
Cell migration requires coordination between the cytoskeleton, adhesion and vesicular traffic. During migration, branched actin networks assemble behind the leading edge, adhesion molecules (integrins) attach to the extracellular matrix and cluster into focal adhesions, and actin stress fibers generate forces by tugging on focal adhesions. Vesicular traffic along microtubules moves proteins to and from the surface and regulates focal adhesion dynamics. Tyrosine kinases such as Src and FAK are recruited to focal adhesions and phosphorylate several focal adhesion proteins. Even though these phosphorylations occur while adhesions are initially forming, the mutation of FAK, Src or the substrates causes excessive focal adhesion growth and inhibits cell migration, suggesting that tyrosine phosphorylation plays a major role in focal adhesion disassembly and emphasizing the importance of focal adhesion turnover for cell movement. Our evidence suggests that ubiquitylation and proteasomal degradation also plays an important role in adhesion dynamics.
Specifically, we found that cell proliferation, transformation and migration are increased when the E3 ubiquitin ligase, CRL5, is inhibited. Further study of CRL5-deficient cells revealed that not only do they migrate more rapidly, but they also have an enlarged leading lamellipodium, increased ruffling at the leading edge, and smaller, more dynamic, focal adhesions. These changes require the tyrosine kinase Src and the focal adhesion protein p130Cas (also known as BCAR1, here referred to as Cas). Phosphorylated Cas is known to recruit proteins that regulate focal adhesion dynamics. We found that phosphorylated Cas is bound by the CRL5 substrate receptor SOCS6. Moreover, inhibiting expression of either SOCS6 or a CRL5 core subunit, Cullin 5, stimulates focal adhesion turnover. Under conditions of decreased CRL5-SOCS6, the amount of phospho-Cas in the focal adhesions at the leading edge increases, presumably because it is being degraded more slowly. We also found that SOCS6 is recruited to focal adhesions by binding to phosphorylated Cas. To test whether phospho-Cas ubiquitylation or degradation is occurring in the cytosol or in focal adhesions, we used optogenetics to restrict SOCS6 access to focal adhesions. Our results suggest that CRL5-SOCS6 only regulates focal adhesion dynamics if it can access Cas in focal adhesions, implying that ubiquitylation, and potentially degradation, of Cas occurs locally and slows focal adhesion disassembly.
Our findings suggest a paradigm in which the local ubiquitylation and degradation of Cas in a particular focal adhesion may slow the disassembly of that focal adhesion. However, we do not yet know how spatially or temporally restricted CRL5-SOCS6 is in its actions on Cas, nor whether other molecules in a focal adhesion are also ubiquitylated.
Current and future plans include:
- Uncovering the time sequence of Cas phosphorylation, SOCS6 translocation, Cas ubiquitylation, and Cas destruction within focal adhesions.
- Identifying the functions of additional SOCS proteins and their substrates in cell migration.
- Dissecting the mechanisms by which CRL5-SOCS regulates cell proliferation and transformation in vitro and development in vivo.


© 2025 Fred Hutchinson Cancer Center, a 501(c)(3) nonprofit organization.